MMV_H4Tracks
Human in the loop 2d cell migration analysis
MMV_H4Tracks (Human4Tracks) is a plugin to use with napari for segmenting and tracking cells, which additionally enables user-friendly manual curation of segmentation and tracks and various options for analyzing and evaluating the results.
We have tested MMV_H4Tracks intensively under Linux and Windows, for Mac there may be problems with parallel computing, which are on our roadmap and will be fixed in a future version.
This napari plugin was generated with Cookiecutter using @napari's cookiecutter-napari-plugin template.
Installation
You can install mmv_h4tracks via pip:
pip install mmv_h4tracks
By default, CPU is used for segmentation computing. We did our best to optimize the CPU computing time, but still recommend GPU computing for better performance. For more detailed instructions on how to install GPU support look here.
Documentation
This plugin was developed to analyze 2D cell migration. It includes the function of segmenting 2D videos using Cellpose (both CPU and GPU implemented) and then tracking them using different automatic tracking algorithms, depending on the use case. For both segmentation and tracking, we have implemented user-friendly options for manual curation after automatic processing. In conjunction with napari's inherent functionalities, our plugin provides the capability to automatically track data and subsequently process the tracks in three different ways based on the reliability of the automated results. Firstly, any potentially existing incorrect tracks can be rectified in a user-friendly manner, thereby maximizing the evaluation of available information. Secondly, unreliable tracks can be selectively deleted, and thirdly, individual tracks can be manually or semi-automatically created for particularly challenging data, ensuring reliable results. In addition, the manually curated results can be compared with the automatic results in order to obtain a score for the quality of the automatic results. In essence, our tool aims to offer a valuable supplement to the existing fully automated tracking tools and a user-friendly means to analyze videos where fully automated tracking has been previously challenging.
Common metrics such as speed, cell size, velocity, etc... can then be extracted, plotted and exported from the tracks obtained in this way. Furthermore, the plugin incorporates a functionality to assess the automatic tracking outcomes using a quality score. Since automated tracking may not be consistently 100% accurate, presenting a quality measure alongside scientific discoveries becomes essential. This supplementary metric offers researchers valuable insights into the dependability of the produced tracking results, fostering informed data interpretation and decision-making in the analysis of cell migration.
More detailed information and instructions on each topic can be found in the following sections.
Get started
To load your raw data, you can simply drag & drop them into napari. Ensure that the 'Image' combobox displays the correct layer afterward, see example:
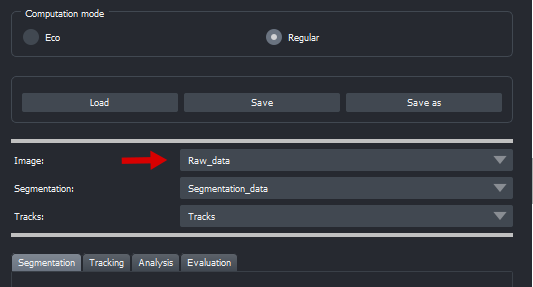
To load your own segmentation, you can do equivalent.
The "save as" button can be used to save the existing layers (raw, segmentation, tracks) in a single .zarr file, which can be loaded again later using the "load" button. The "save" button overwrites the loaded .zarr file.
The computation mode is used to set how many of the available CPU cores (40% or 80%) are to be used for computing the CPU segmentation and tracking and therefore has a direct impact on performance.
Segmentation
For segmentation, we use the state of the art instance segmentation method Cellpose. We provide a model that we trained and has proven successful for our application (see more information). Note: Due to computational constraints, we do not employ Cellpose-SAM here.
Automatic instance segmentation
To start automatic segmentation, a model must first be selected. Automatic segmentation can then be started via "Run Segmentation". The "Preview" option offers the possibility of segmenting the first 5 frames first in order to obtain an estimate of the expected results, as the computation - depending on the data and hardware - can be time-consuming.
Add custom models
The plugin supports adding custom Cellpose models. To do so, simply click on "Add custom Cellpose model", enter a name to be displayed, select the model path and pass the required parameters. Click here for more information about the parameters.
To train your own Cellpose model, this might be helpful.
Train new models
The plugin also allows users to train new Cellpose models. For this, expand “Train new Cellpose model”, select the frames to use for training, provide a model name, and click “Train model.” Once training finishes successfully, a popup prompts the user to choose the path where the model should be saved.
By default, training runs for 200 epochs. If the “Longer Training” option is enabled, the number of training epochs is increased to 1000.
Models trained within the plugin remain available after restarting it. If the plugin is reinstalled or the model needs to be used on another computer, it can simply be re-added as a custom model.
Manual curation
We provide different options to correct the automatic segmentation:
Remove cell- Click on a cell to remove it. Be aware that removing a cell will split the track the cell belongs to, potentially affecting subsequent tracking.Next free ID- Loads the next free label ID, then a false negative cell can be manually annotated using the paint mode.Select ID- Click on a cell to load its ID, then this cell can be corrected manually using the paint mode.Merge cell- Click on 2 different fragments of the same cell to harmonize their ID. Note: This has no effect on the annotation itself.Separate- Click on a cell to assign a new ID to it.
Tracking
The plugin supports both coordinate-based (LAP) and overlap-based tracking. Overlap-based tracking requires more computation, but can also be used in particularly complicated data for individual cells. In our experience, coordinate-based tracking has proven itself in cases with reliable segmentation. Overlap-based tracking serves as a useful complement in cases where the segmentation is not of sufficient quality.
If necessary, overlap-based tracking can also be used for single cells. To do this, simply click on the cell after clicking the button.
Manual curation
To correct tracks, the plugin allows you to link or unlink them. For both options, first click on the corresponding button and then on the cell in the respective frame. The action must then be confirmed using the previously clicked button, which now displays "confirm".
To unlink, all you need to do is click on the cell in the first and last frame. So if the cell is tracked from frame 1-100 and the track between frames 1-10 is to be deleted, it is sufficient to click on the cell in frames 1 and 10. If the track is to be deleted between frames 40-60, it is sufficient to click in frames 40 and 60. In this scenario, the rest of the track is then split, i.e. once into a track from frame 1-40 and once into a track from frame 60-100.
In contrast, to link cells, the corresponding cell in each frame must be clicked. This must be done for all frames, so the track must be gapless.
Visualize & filter tracks
The displayed tracks can be filtered by entering specific track IDs. Click on the "Show all tracks" button to display all tracks again.
Individual tracks can be deleted using the delete function. Note: These are permanently deleted and cannot be restored without re-tracking. In addition, all displayed tracks can be deleted.
Analysis
The plugin supports the calculation of various metrics, which can be divided into two categories: migration-based (such as speed, direction, ...) and shape-based (such as size, eccentricity, ...). Through the use of these metrics, a comprehensive understanding of the available data can be obtained.
All these metrics can be exported to a .csv file. In addition, the tracks can be filtered with a movement minimum (in pixels) and a minimum track length (in frames). Note: All existing tracks are exported in any case, but their results are presented separately.
The plugin offers the option of filtering the existing tracks according to the metrics. To do this, the corresponding metric can be selected in the plot area and a scatter plot of the data points will be generated using the plot button. Individual data points (/tracks) that are to be displayed can be circled with the mouse and all tracks that are not circled will be hidden. Note: No tracks are deleted in this process. Hiding tracks triggers the filter function in the tracking section, the "Show all tracks" button can display all tracks again as described above.
Evaluation
To be aware of the accuracy of your automatic tracking and segmentation results, we have implemented an option to evaluate your automatic results. Evaluation is always carried out against the latest results of automatic segmentation and automatic tracking or previously created results loaded via the plugin's own load function. We may implement the option to evaluate external segmentations in the future, but for now you can use save and load as a workaround.
To evaluate results, at least two consecutive frames must be manually corrected first. The plugin saves the previously mentioned automatic or loaded results in the background, so no activation via button or similar is necessary before manual correction.
The range of frames to be evaluated can be set, for which the results for segmentation and tracking can be calculated independently of each other.
Segmentation evaluation
In order to evaluate the segmentation results, a segmentation must first be loaded via the plugin's load function (drag & drop via napari is not sufficient) or computed within the plugin. This can then be corrected manually. For IoU, Dice and Average Precision 50 scores are then calculated for the frames specified by the user. These results are not exported automatically and must therefore be noted down by users themselves.
Tracking evaluation
As for the evaluation of the segmentation, tracking results loaded via the plugin or obtained within the plugin are required. At least 2 consecutive frames must be corrected manually so that a score can be calculated for the quality of the tracking results. More information can be found here.
Assistant
The assistant tab serves to facilitate the identification of errors within segmentation and tracking. It is divided into filters and segmentation adaptation.
Recommended filter strategy:
- "Show noteworthy tracks" to discover tracks that are not close to the edge and emerge or disappear after the movie starts. Tracks must be gapless, and resulting hits can be indications of errors.
- "Show small cells" to double-check if there is any noise/pollution segmented.
- "Show untracked cells" to identify untracked segmented instances.
The other filters can provide additional support.
The segmentation adaptation functions supplement useful functions with respect to segmentation. "Align segmentation IDs" adapts the label IDs with regard to the tracking IDs (the label IDs are then no longer arbitrary). "Relabel cells" ensures that the labels within a frame are unique, which helps eliminate accidental errors during manual segmentation correction.
Hotkeys
Here's an overview of the hotkeys. All of them can also be found in the corresponding tooltips.
W- Load next free segmentation IDG- Overlap-based single cell trackingH- Separate cellsQ- Select cell ID
Development plan
We will continue to develop the plugin and implement new features in the future. Some of our plans in arbitrary order:
- Feedback (progress bar) for computationally intensive functions
- Support of lineages
Support training custom Cellpose models within the plugin- Model optimization to further optimize segmentation computation
- Support evaluation of external segmentations
- Improve robustness of Mac computing
- Batch processing
- Add support for OME-Zarr
- Add support for multiscale data
- Add single-cell tracking view (action cam)
If you have a feature request, please file an issue.
Resources
The following resources may be of interest:
Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the BSD-3 license, "mmv_h4tracks" is free and open source software
Issues
If you encounter any problems, please file an issue along with a detailed description.
Version:
- 1.3.0
Last updated:
- 2026-03-27
First released:
- 2024-03-22
License:
- Copyright (c) 2022, lennart ko...
Operating system:
- Information not submitted
Requirements:
- numpy
- npe2
- napari-plugin-engine>=0.1.4
- napari
- zarr>=3.0.8
- cellpose<4
- tqdm>=4.0
- matplotlib
- bioio
- scipy>=1.11.0
- ome-zarr>=0.12.0
- napari-ome-zarr
- napari[all]; extra == "all"
- pytest; extra == "all"
- pytest-qt; extra == "all"
- pre-commit; extra == "all"
- bioio-tifffile; extra == "all"





